Overview
In today's Information Age, the human world and especially science increasingly revolve around huge amounts of data. It is very important for the success of our project to create a robust data handling system that addresses many types of data. Our initial data concerns will be to address:
- Climate change simulations
- Plant communities modeling
- Animal species observations
- Challenges for data software design
- People need data too
1. Climate Change Simulations
Global climate models (GCMs) generate mountains of 4-dimensional data that record the behavior of numerous state variables across multiple decades of time. Fortunately, we are usually interested in just the mean monthly values for temperature, precipitation, and relative humidity, averaged over the equilibrium period of each simulation.
We are already using climate change projections published by the IPCC from the SRES-A1B intermediate scenario for plant community studies. Both climate and vegetation fields have been stored in NetCDF files, a format developed by UCAR's UniData division, which is currently based on the HDF5 format, created by NASA for storing huge amounts of remote sensing data. Unlike most database management systems, NetCDF is designed to accommodate large multidimensional arrays. Please read the Data Challenges section below for a more in-depth look at these contrasting approaches to data.
2. Plant Communities Modeling
Our Equilibrium Vegetation Ecology model (EVE) transforms monthly climatologies into plant community structure as defined by the areal coverage of any of 110 plant life forms. Like the GCMS, EVE generates large multi-dimensional arrays as its output.
As stated above, results from plant community simulations are currently being stored in NetCDF files. This will remain the data format of choice for our large gridded (raster) datasets, until it has been determined that the new raster capabilities of one of the spatial database systems is really ready and worth the investment of time required to switch over to that option.
3. Animal Species Observations
Our current animal studies involve making observations of populations at sites around the world. In contrast to the gridded data generated by climate and vegetation models, our animal data describe population levels observed at "random" geographic points.
Our animal (lizard) population data are currently being recorded for individual sites in tables in Excel spreadsheets. These data can easily be imported into a standard database for storage and for serving up to our web mapping system.
4. Challenges for Data Software Design
Our gridded arrays are raster data while our site observations behave as vector data. As these two data worlds have mostly evolved on separate tracks, harmonizing them in a single data solution is challenging. A summary of this challenge follows.
The NetCDF file format is one of the premier choices for storing gridded data, such as those generated by global climate models (GCMs). NetCDF easily supports large multidimensional arrays. This quote is taken directly from the Unidata NetCDF manual:
NetCDF Is Not a Database Management System
Why not use an existing database management system for storing array-oriented data? Relational database software is not suitable for the kinds of data access supported by the netCDF interface.
First, existing database systems that support the relational model do not support multidimensional objects (arrays) as a basic unit of data access. Representing arrays as relations makes some useful kinds of data access awkward and provides little support for the abstractions of multidimensional data and coordinate systems. A quite different data model is needed for array-oriented data to facilitate its retrieval, modification, mathematical manipulation and visualization.
Related to this is a second problem with general-purpose database systems: their poor performance on large arrays. Collections of satellite images, scientific model outputs and long-term global weather observations are beyond the capabilities of most database systems to organize and index for efficient retrieval.
Finally, general-purpose database systems provide, at significant cost in terms of both resources and access performance, many facilities that are not needed in the analysis, management, and display of array-oriented data. For example, elaborate update facilities, audit trails, report formatting, and mechanisms designed for transaction-processing are unnecessary for most scientific applications.
5. People need data too
Community science must deal not only with its collective scientific data, but also must keep track of the scientists that compose the community.
In parallel to developing a comprehensive data system for handling the biodiversity data, we must also implement a data system for keeping track of our community members. This is precisely the sort of thing that traditional table-based databases were designed to accomplish and so we will probably create a PostgreSQL database connected to the website for this purpose.
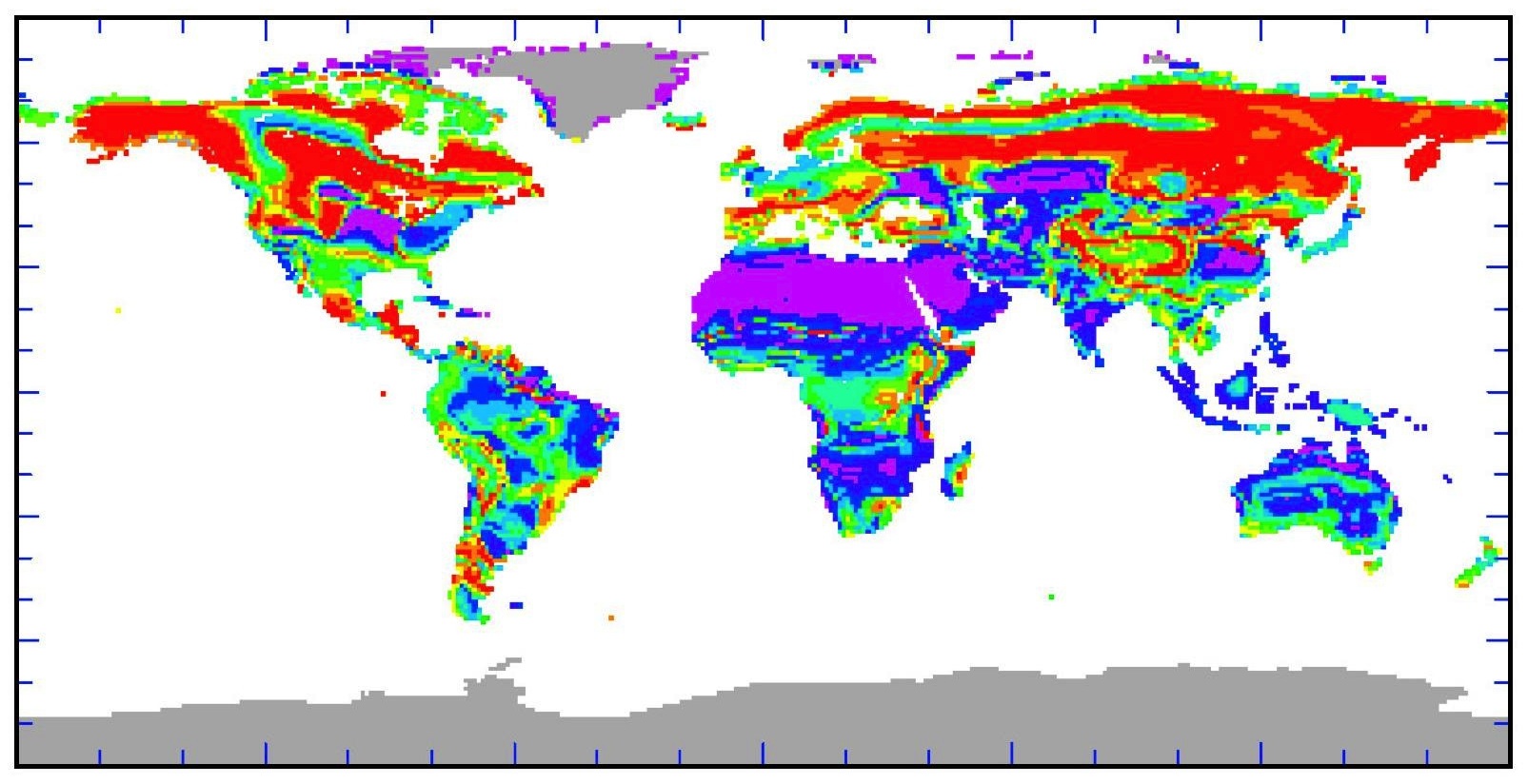
Ecological sensitivity map